WinBUGS Tutorial
WinBUGs
WinBUGS (Windows version of Bayesian inference Using Gibbs Sampling) is a foundational tool in Bayesian statistics. While newer engines like JAGS or Stan are common now, WinBUGS remains a “gold standard” for learning because its GUI and “DoodleBUGS” help visualize how parameters interact.
Notes
Windows Installation: Installing WinBUGS on a Windows OS is straightforward and requires no additional configuration.
macOS Installation: Recent Mac models (M2/M3 chips) operate exclusively on 64-bit architecture, which is incompatible with the 32-bit requirements of WinBUGS. To bypass this, UB students, faculty, and staff are encouraged to use the UB Virtual Public Site. This service provides a virtual Windows environment where WinBUGS can be accessed and run easily. Alternatively, users may consider installing JAGS, a cross-platform equivalent that natively supports Windows, macOS, and Linux.
Install
- Download zip file using via this link.
- Extract the zip file and open the Application named
WinBUGS
Example
Scenario: The “Eight Schools” example. We want to estimate the effect of coaching in 8 different schools, assuming schools are similar but not identical (they “borrow strength” from each other).
In a hierarchical model, we assume that the parameters of interest are not fixed, but are instead drawn from a common “population” distribution. We structure this in three levels:
Level 1: The Likelihood (Data Level)
Each school $j$ has an observed effect $y_j$ with a known standard error $\sigma_j$:
\[y_j \mid \theta_j \sim N(\theta_j, \sigma_j^2), \quad j = 1, \dots, 8\]Level 2: The Population (Structural Level)
We assume the true effects $\theta_j$ are not independent. They are “linked” by a shared distribution:
\[\theta_j \mid \mu, \tau \sim N(\mu, \tau^2)\]$\mu$: The “grand mean” (the average coaching effect across all schools).
$\tau$: The “between-school” variability.
Level 3: The Hyper-priors
Since we are Bayesian, we must place priors on the parameters of our population distribution:
\[\mu \sim \text{Non-informative Prior (e.g., } N(0, 10^6))\] \[\tau \sim \text{Weakly Informative Prior (e.g., } \Gamma(0.001, 0.001))\]Note:
- Stochastic Nodes (~): These represent uncertainty. They MUST be initialized. This includes your parameters and your data (though data is initialized by the “Load Data” step).
- Logical Nodes (<-): These are deterministic links. They are calculated automatically from other nodes, so you NEVER provide initial values for them.
model {
for (j in 1:J) {
# Likelihood for each school
y[j] ~ dnorm(theta[j], tau.y[j])
# School-specific effects drawn from a "Population" distribution
theta[j] ~ dnorm(mu.theta, tau.theta)
# Standard error for each school is usually known data
tau.y[j] <- 1 / (sigma.y[j] * sigma.y[j])
}
# Hyper-priors (The "Global" parameters)
mu.theta ~ dnorm(0, 0.0001)
tau.theta ~ dgamma(0.01, 0.01)
# Global Standard Deviation
sd.theta <- 1 / sqrt(tau.theta)
}
# DATA
list(J = 8,
y = c(28.39, 7.94, -2.75, 6.82, -0.64, 0.63, 18.01, 12.16),
sigma.y = c(14.9, 10.2, 16.3, 11.0, 9.4, 11.4, 10.4, 17.6))
# INITS
list(mu.theta = 0, tau.theta = 1, theta = c(0,0,0,0,0,0,0,0))
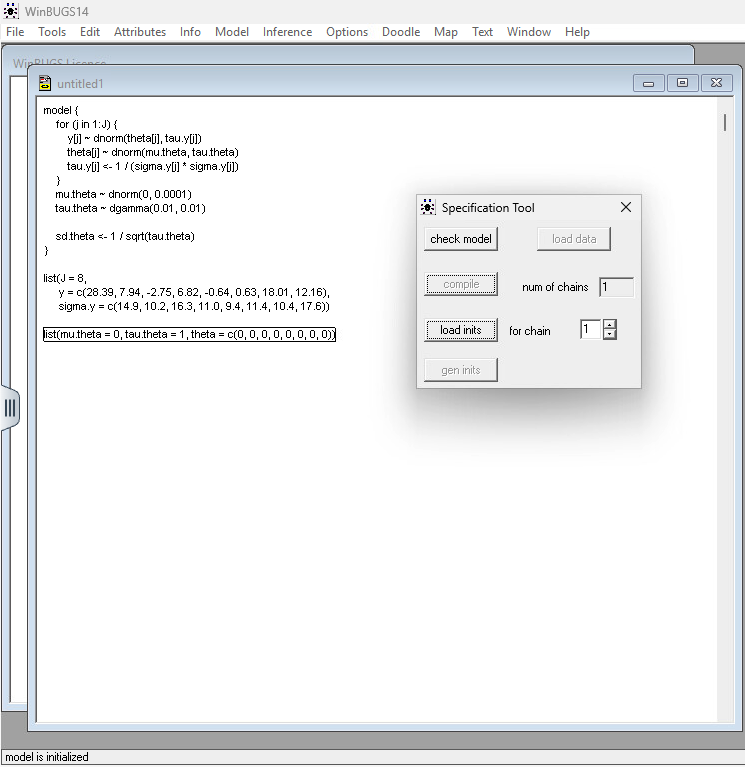
Step 1: Use File > New to open a new document Step 2: Copy the code above inside the workspace Step 3: Click Model > Specification
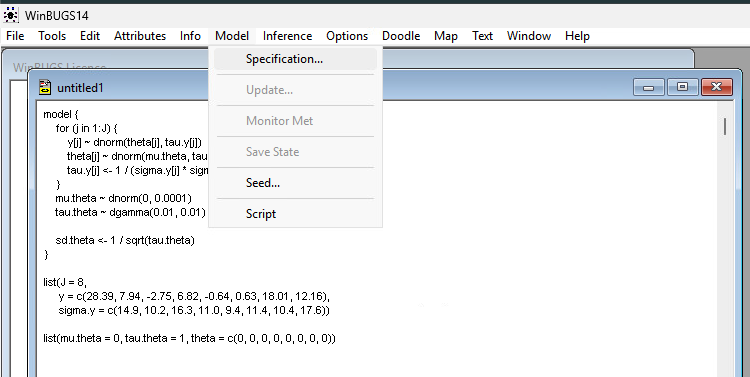
Step 4: Check Model: Highlight the word model in your code and click “Check Model.” The status bar (bottom left) should say “model is syntactically correct.”
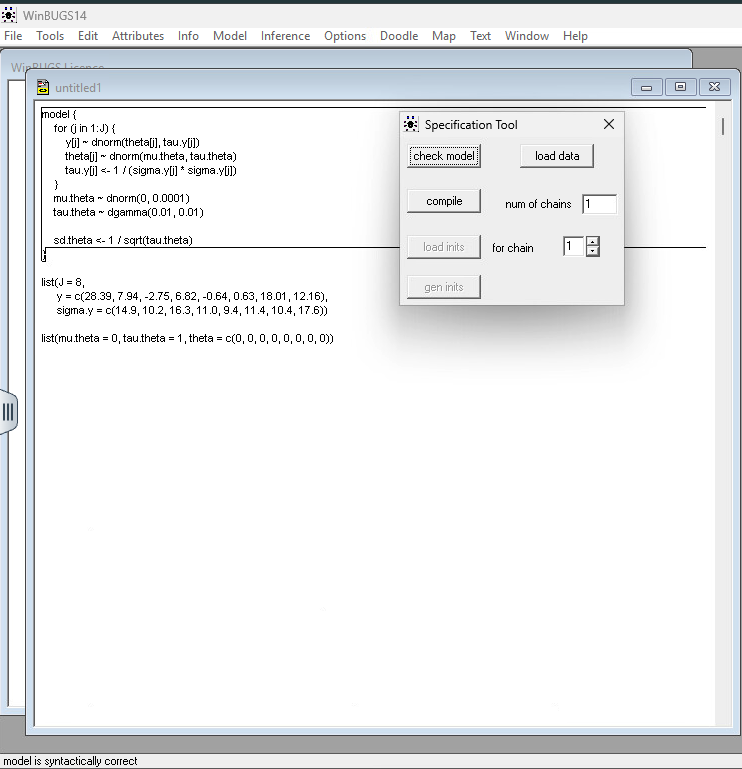
Step 5: Load Data: Highlight the word list in your data block and click “Load Data.”
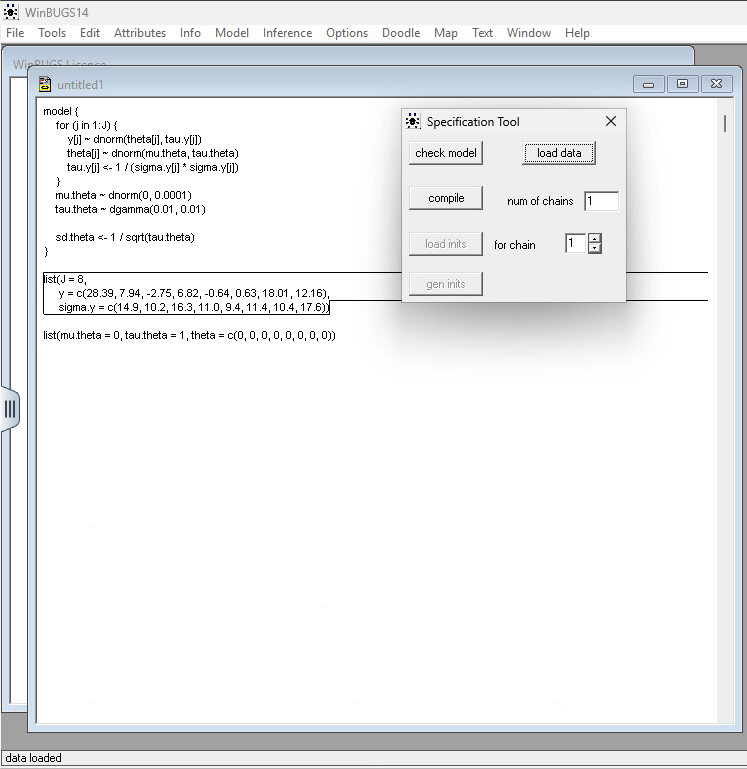
Step 6: Compile: Click “Compile.” This builds the internal mathematical graph. Step 7: Load Inits: Highlight the list in your initial values block and click “Load Inits.” If you’re lazy, you can click “Gen Inits,” but providing your own is safer for complex models.
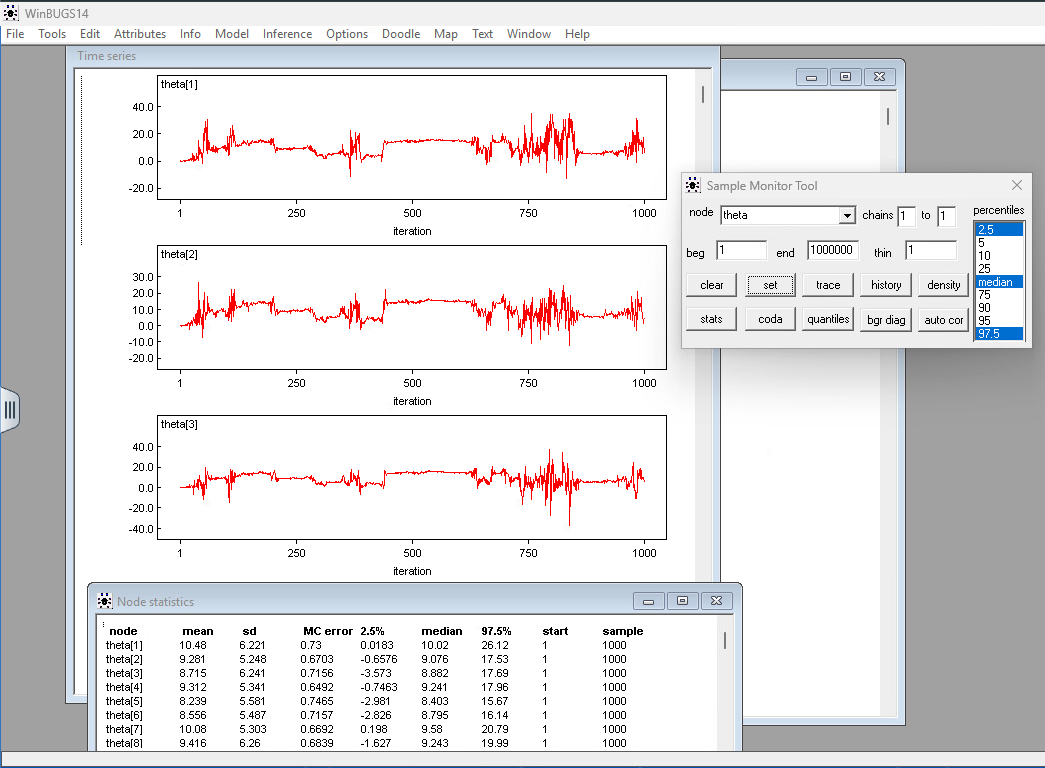
To view trace plots for an array of variables like $\theta_j$ (the individual effects for your 8 schools):
Step 1: Open Inference > Samples
Step 2: In the node field, type simply theta
Step 3: Click Set. This tells WinBUGS to start recording the values for every index of that array. Step 4: Open Model > Update. Run update (e.g., 1,000 or 10,000 iterations).
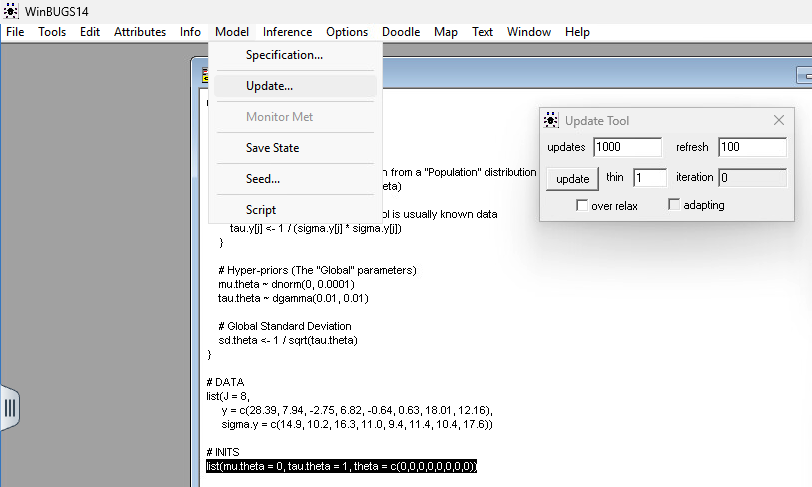
Step 5: Go back to the Sample Monitor Tool, type theta in the node field again, and click History to view a graphics window showing the sample trace or stats to view the summary statistics for the variable, pooling over the chains selected.
DoodleBUGS
DoodleBUGS is essentially a “drag-and-drop” editor for building Directed Acyclic Graphs (DAGs). For your presentation, demonstrating how a visual graph translates into statistical code is a powerful way to show you understand the hierarchy.
Opening the Workspace
Step 1: Go to Model > Doodle. A blank white window will appear.
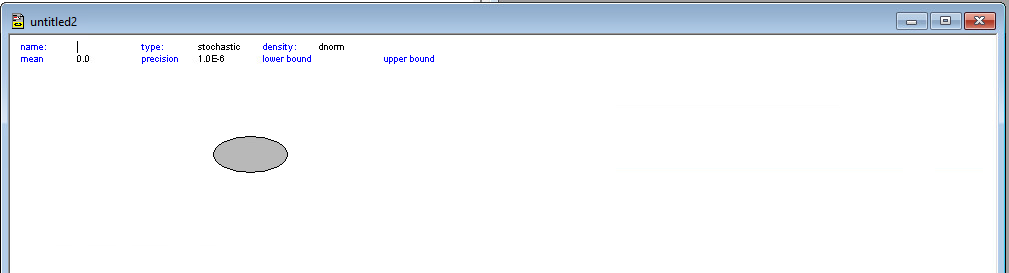
Step 2: Your cursor is now a “tool.” Notice the toolbar at the top of the Doodle window: - Oval: Stochastic Node ($\sim$) - Rectangle: Logical Node ($\leftarrow$) - Plate (Stacked Rectangles): For loops (the $j$ index) - Arrow: Dependencies
Building the Hierarchy Step-by-Step
- Step A: The “School” Plate: Since you have 8 schools, you need a Plate to represent the loop
for (j in 1:J).- Click the Plate tool (
ctrl+ click an empty region) and draw a large box in the middle of the screen. - Double-click the edge of the plate. Name it
jand set the range from1to8.
- Click the Plate tool (
- Step B: The Individual Effects ($\theta_j$)
- Select the Oval tool (Stochastic).
- Click inside the plate to create a node. Double-click it and name it
theta. - Because it’s inside the plate, WinBUGS automatically knows it is an array:
theta[j].
- Step C: The Observed Data ($y_j$)
- Create another Oval inside the plate. Name it
y. - Use the Arrow tool to draw a line from
thetatoy(Selectythenctrl+ clicktheta). This tells the model that $y$ depends on $\theta$.
- Create another Oval inside the plate. Name it
- Step D: The Hyper-Parameters ($\mu$ and $\tau$): These parameters are shared by all schools, so they must live outside the plate.
- Create two Ovals outside the plate. Name them
mu.thetaandtau.theta. - Draw arrows from both
mu.thetaandtau.thetainto the theta node inside the plate.
- Create two Ovals outside the plate. Name them
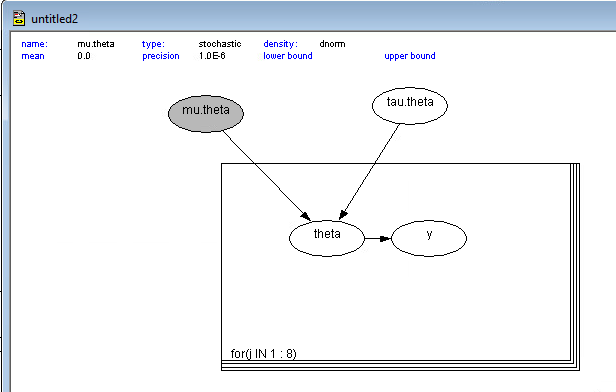
Defining the Distributions
Once the “skeleton” is drawn, you must define the math. Double-click each node:
- For $y$: Set the density to
dnorm. In the parameters, link it tothetaand its own precision (or a constant if $\sigma$ is known). - For $\theta$: Set the density to
dnorm. Link the mean tomu.thetaand the precision totau.theta. - For $\mu$ and $\tau$: Set these as your priors (e.g., dnorm(0, 0.001)).
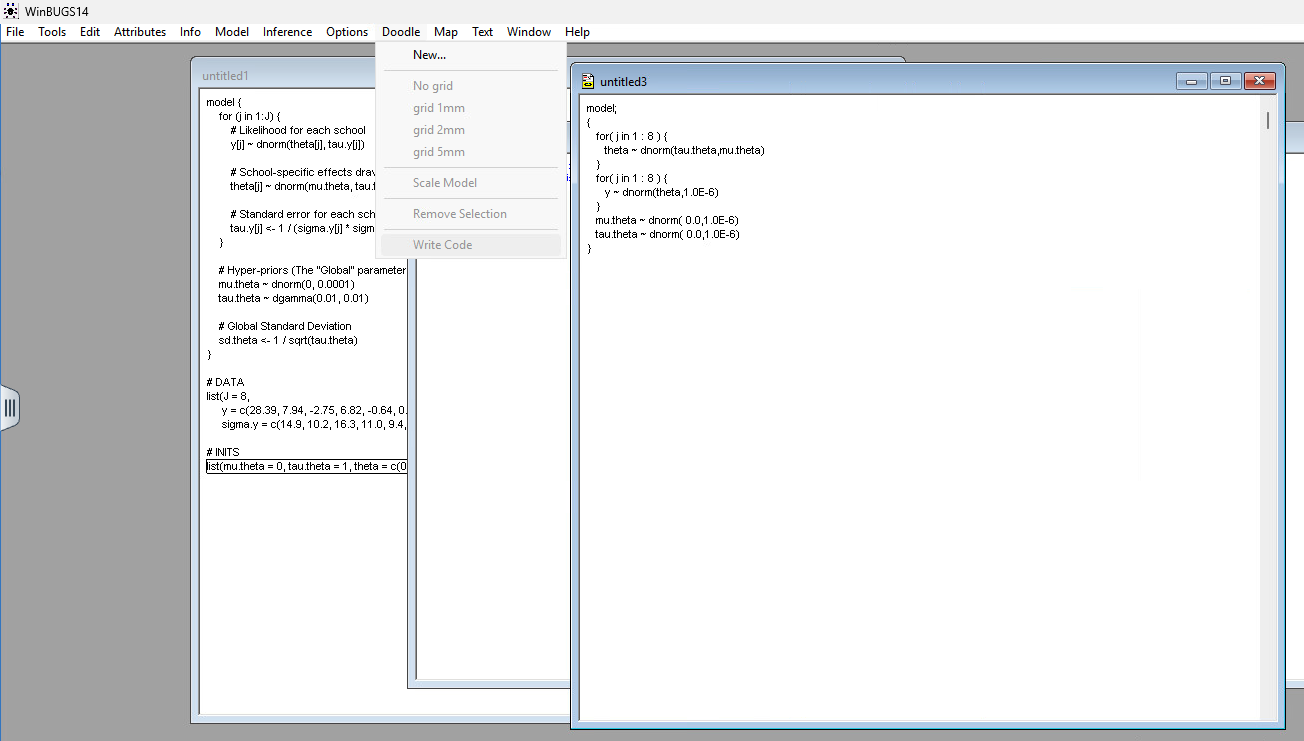
Write Code
Once your Doodle is finished, go to the Doodle menu at the top of the screen. Click Write Code. WinBUGS will instantly open a new text window with the exact BUGS code we used in the above example.
R2WinBUGS
R2WinBUGS is a classic R package that acts as a bridge between the R environment and the WinBUGS software. It allows you to write your Bayesian models in the BUGS language but manage the data, execution, and output entirely within R.
Step 1: Install R2WinBUGS Package
install.packages("R2WinBUGS")
Step 2: Run the above example by passing the same WinBUGs code directly into R2WinBUGS package:
library(R2WinBUGS)
# Create the model file
cat("
model {
for (j in 1:J) {
y[j] ~ dnorm(theta[j], tau.y[j])
theta[j] ~ dnorm(mu.theta, tau.theta)
tau.y[j] <- 1 / (sigma.y[j] * sigma.y[j])
}
mu.theta ~ dnorm(0, 0.0001)
tau.theta ~ dgamma(0.01, 0.01)
sd.theta <- 1 / sqrt(tau.theta)
}
", file = "8schools_model.txt")
schools_data <- list(
J = 8,
y = c(28.39, 7.94, -2.75, 6.82, -0.64, 0.63, 18.01, 12.16),
sigma.y = c(14.9, 10.2, 16.3, 11.0, 9.4, 11.4, 10.4, 17.6)
)
schools_inits <- function() {
list(mu.theta = 0, tau.theta = 1, theta = rep(0, 8))
}
params <- c("mu.theta", "sd.theta", "theta")
schools_sim <- bugs(
data = schools_data,
inits = schools_inits,
parameters.to.save = params,
model.file = "8schools_model.txt",
n.chains = 3,
n.iter = 5000,
n.burnin = 1000,
bugs.directory = "C:/Users/kevinyen/Downloads/winbugs143_unrestricted/winbugs14_full_patched/WinBUGS14",
debug = TRUE # Set to TRUE to see the WinBUGS window and debug errors
)
print(schools_sim)
> print(schools_sim)
Inference for Bugs model at "8schools_model.txt", fit using WinBUGS,
4 chains, each with 2000 iterations (first 1000 discarded), n.thin = 4
n.sims = 1000 iterations saved
mean sd 2.5% 25% 50% 75% 97.5% Rhat n.eff
mu.theta 7.8 3.9 0.9 5.0 7.6 10.2 16.3 1.1 43
sd.theta 1.8 2.4 0.1 0.3 0.9 2.2 8.6 1.0 79
theta[1] 8.4 4.8 0.5 5.3 7.9 10.8 18.6 1.0 75
theta[2] 7.8 4.4 -0.6 4.9 7.4 10.3 16.9 1.1 51
theta[3] 7.4 4.5 -1.5 4.7 7.1 10.1 16.5 1.1 51
theta[4] 7.7 4.4 -1.4 4.9 7.5 10.2 16.8 1.1 46
theta[5] 7.3 4.5 -2.2 4.6 7.1 9.9 16.5 1.1 56
theta[6] 7.4 4.3 -0.7 4.5 7.2 10.0 16.5 1.1 44
theta[7] 8.2 4.7 -0.1 5.1 7.8 10.7 18.0 1.1 56
theta[8] 7.9 4.7 -0.8 5.0 7.8 10.4 17.2 1.1 53
deviance 60.3 1.5 58.2 59.5 59.9 60.6 64.1 1.0 910
For each parameter, n.eff is a crude measure of effective sample size,
and Rhat is the potential scale reduction factor (at convergence, Rhat=1).
DIC info (using the rule, pD = Dbar-Dhat)
pD = 1.2 and DIC = 61.5
DIC is an estimate of expected predictive error (lower deviance is better).
JAGs
JAGS stands for Just Another Gibbs Sampler. It is a cross-platform program designed for the analysis of Bayesian hierarchical models using Markov Chain Monte Carlo simulation.
Developed by Martyn Plummer, it was built to be a more portable and extensible alternative to the original BUGS engines (WinBUGS and OpenBUGS). In modern biostatistics, it is often the “next step” after learning the basics of WinBUGS.
Install JAGs on Windows
https://sourceforge.net/projects/mcmc-jags/files/JAGS/4.x/Windows/
install.packages("rjags")
library(rjags)
Install JAGs on Mac
First install the JAGs
brew install jags
Directly install in this way
install.packages("rjags", type = "source",
configure.args = "--with-jags-include=/opt/homebrew/include/JAGS --with-jags-lib=/opt/homebrew/lib")
You should see these long messages since it directly install package directly by source build
> install.packages("rjags", type = "source",
+ configure.args = "--with-jags-include=/opt/homebrew/include/JAGS --with-jags-lib=/opt/homebrew/lib")
trying URL 'https://cran.rstudio.com/src/contrib/rjags_4-17.tar.gz'
Content type 'application/x-gzip' length 76394 bytes (74 KB)
==================================================
downloaded 74 KB
* installing *source* package ‘rjags’ ...
** this is package ‘rjags’ version ‘4-17’
** package ‘rjags’ successfully unpacked and MD5 sums checked
** using staged installation
configure: WARNING: unrecognized options: --with-jags-include, --with-jags-lib
checking for pkg-config... no
configure: Attempting legacy configuration of rjags
checking for jags... /opt/homebrew/bin/jags
checking whether the C++ compiler works... yes
checking for C++ compiler default output file name... a.out
checking for suffix of executables...
checking whether we are cross compiling... no
checking for suffix of object files... o
checking whether the compiler supports GNU C++... yes
checking whether clang++ -arch arm64 -std=gnu++17 accepts -g... yes
checking for clang++ -arch arm64 -std=gnu++17 option to enable C++11 features... none needed
checking for stdio.h... yes
checking for stdlib.h... yes
checking for string.h... yes
checking for inttypes.h... yes
checking for stdint.h... yes
checking for strings.h... yes
checking for sys/stat.h... yes
checking for sys/types.h... yes
checking for unistd.h... yes
checking for Console.h... yes
configure: Compile flags are -I/opt/homebrew/include/JAGS
configure: Link flags are -L/opt/homebrew/lib -ljags
checking for gcc... clang -arch arm64 -std=gnu2x
checking whether the compiler supports GNU C... yes
checking whether clang -arch arm64 -std=gnu2x accepts -g... yes
checking for clang -arch arm64 -std=gnu2x option to enable C11 features... none needed
checking for jags_version in -ljags... yes
checking version of JAGS library... OK
configure: creating ./config.status
config.status: creating src/Makevars
configure: WARNING: unrecognized options: --with-jags-include, --with-jags-lib
configure: creating ./config.status
config.status: creating src/Makevars
config.status: creating R/unix/zzz.R
configure: WARNING: unrecognized options: --with-jags-include, --with-jags-lib
** libs
using C compiler: ‘Apple clang version 21.0.0 (clang-2100.0.123.102)’
using C++ compiler: ‘Apple clang version 21.0.0 (clang-2100.0.123.102)’
using SDK: ‘MacOSX26.4.sdk’
clang -arch arm64 -std=gnu2x -I"/Library/Frameworks/R.framework/Resources/include" -DNDEBUG -I/opt/homebrew/include/JAGS -I/opt/R/arm64/include -fPIC -falign-functions=64 -Wall -g -O2 -c init.c -o init.o
clang++ -arch arm64 -std=gnu++17 -I"/Library/Frameworks/R.framework/Resources/include" -DNDEBUG -I/opt/homebrew/include/JAGS -I/opt/R/arm64/include -fPIC -falign-functions=64 -Wall -g -O2 -c jags.cc -o jags.o
clang++ -arch arm64 -std=gnu++17 -I"/Library/Frameworks/R.framework/Resources/include" -DNDEBUG -I/opt/homebrew/include/JAGS -I/opt/R/arm64/include -fPIC -falign-functions=64 -Wall -g -O2 -c parallel.cc -o parallel.o
clang++ -arch arm64 -std=gnu++17 -dynamiclib -Wl,-headerpad_max_install_names -undefined dynamic_lookup -L/Library/Frameworks/R.framework/Resources/lib -L/opt/R/arm64/lib -o rjags.so init.o jags.o parallel.o -L/opt/homebrew/lib -ljags -F/Library/Frameworks/R.framework/.. -framework R
installing to /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/library/00LOCK-rjags/00new/rjags/libs
** R
** data
** byte-compile and prepare package for lazy loading
** help
*** installing help indices
** building package indices
** testing if installed package can be loaded from temporary location
** checking absolute paths in shared objects and dynamic libraries
** testing if installed package can be loaded from final location
** testing if installed package keeps a record of temporary installation path
* DONE (rjags)
The downloaded source packages are in
‘/private/var/folders/3s/rg9c_f490gg1t18_0jftcw1m0000gn/T/RtmpN0lS1t/downloaded_packages’
In order to make sure what you do so far is correct, load the rjags library in R:
> library(rjags)
Loading required package: coda
Linked to JAGS 4.3.2
Loaded modules: basemod,bugs
Bug Fix
Issue comes when directly install rjags in this way
install.packages("rjags")
Before fixing, need to remove the files installed before
sudo rm -rf /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/library/rjags
Demo
library(rjags)
# Create the model file
cat("
model {
for (j in 1:J) {
y[j] ~ dnorm(theta[j], tau.y[j])
theta[j] ~ dnorm(mu.theta, tau.theta)
tau.y[j] <- 1 / (sigma.y[j] * sigma.y[j])
}
mu.theta ~ dnorm(0, 0.0001)
tau.theta ~ dgamma(0.01, 0.01)
sd.theta <- 1 / sqrt(tau.theta)
}
", file = "8schools_model.txt")
# Data
jags_data <- list(
J = 8,
y = c(28.39, 7.94, -2.75, 6.82, -0.64, 0.63, 18.01, 12.16),
sigma.y = c(14.9, 10.2, 16.3, 11.0, 9.4, 11.4, 10.4, 17.6)
)
# Initial Values
jags_inits <- list(
list(mu.theta = 0, tau.theta = 1, theta = rep(0, 8)),
list(mu.theta = 10, tau.theta = 0.5, theta = rep(5, 8)) # Chain 2
)
# Initialize / Compile the Model
model_run <- jags.model(
file = "8schools_model.txt",
data = jags_data,
inits = jags_inits,
n.chains = 2
)
# Update Model
update(model_run, n.iter = 1000)
samples <- coda.samples(
model = model_run,
variable.names = c("mu.theta", "sd.theta", "theta"),
n.iter = 5000
)
summary(samples)
> summary(samples)
Iterations = 1001:6000
Thinning interval = 1
Number of chains = 2
Sample size per chain = 5000
1. Empirical mean and standard deviation for each variable,
plus standard error of the mean:
Mean SD Naive SE Time-series SE
mu.theta 8.452 4.397 0.04397 0.3167
sd.theta 2.564 3.311 0.03311 0.1796
theta[1] 9.476 5.711 0.05711 0.3442
theta[2] 8.373 4.958 0.04958 0.3050
theta[3] 7.963 5.513 0.05513 0.3141
theta[4] 8.316 5.015 0.05015 0.3259
theta[5] 7.550 5.111 0.05111 0.3191
theta[6] 7.850 5.139 0.05139 0.3032
theta[7] 9.305 5.172 0.05172 0.3274
theta[8] 8.640 5.496 0.05496 0.3496
2. Quantiles for each variable:
2.5% 25% 50% 75% 97.5%
mu.theta 0.124808 5.2752 8.526 11.603 16.77
sd.theta 0.097733 0.4243 1.280 3.372 11.96
theta[1] -0.008631 5.5878 9.128 12.568 21.97
theta[2] -1.455128 5.0925 8.459 11.668 17.48
theta[3] -3.567675 4.7376 8.102 11.549 17.70
theta[4] -1.616398 5.0578 8.388 11.645 17.73
theta[5] -3.658888 4.4647 7.675 11.123 16.70
theta[6] -2.798620 4.7096 7.965 11.394 17.13
theta[7] 0.084474 5.6810 9.085 12.396 20.28
theta[8] -1.669447 5.2056 8.614 11.960 19.09
plot(samples)
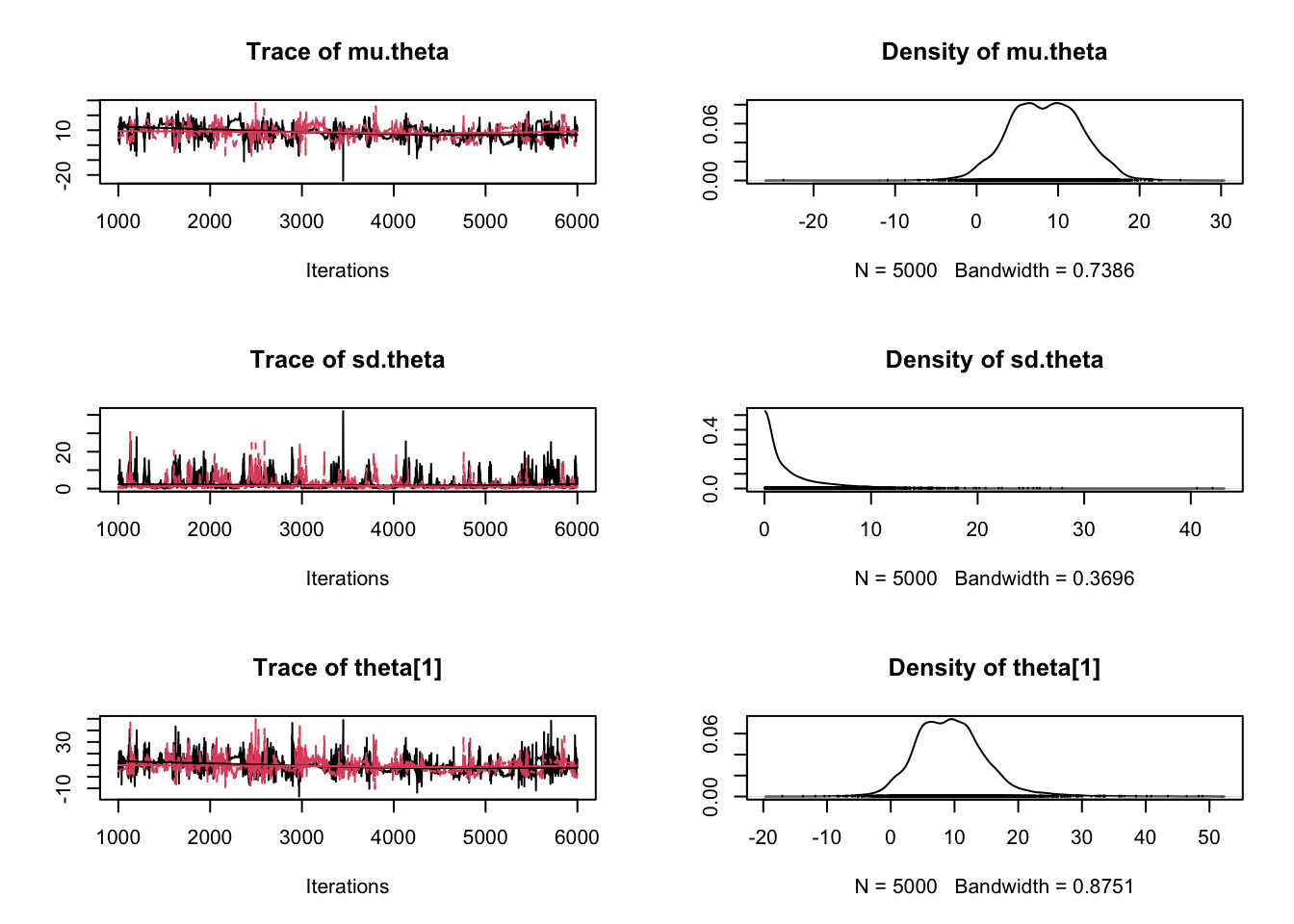
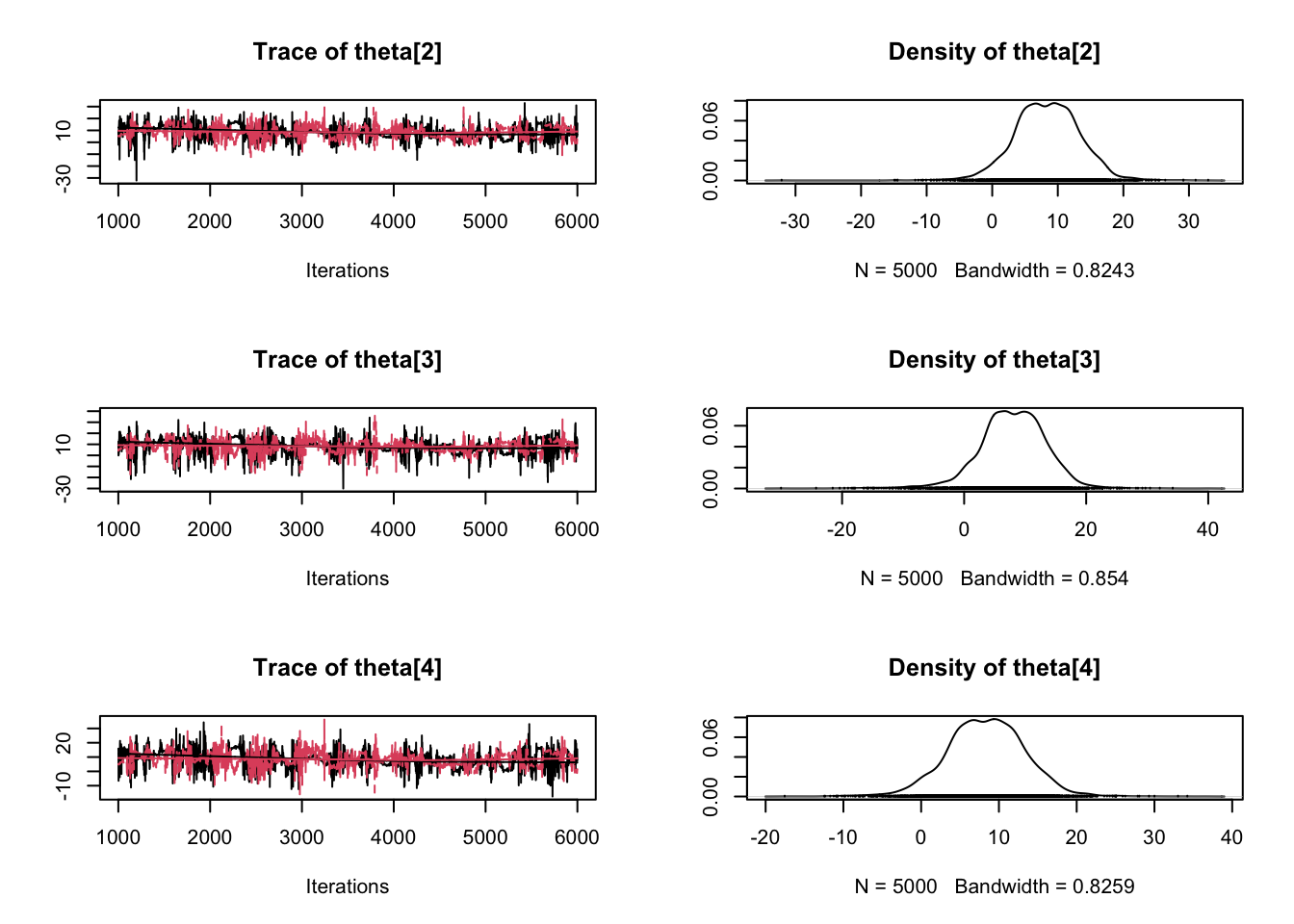
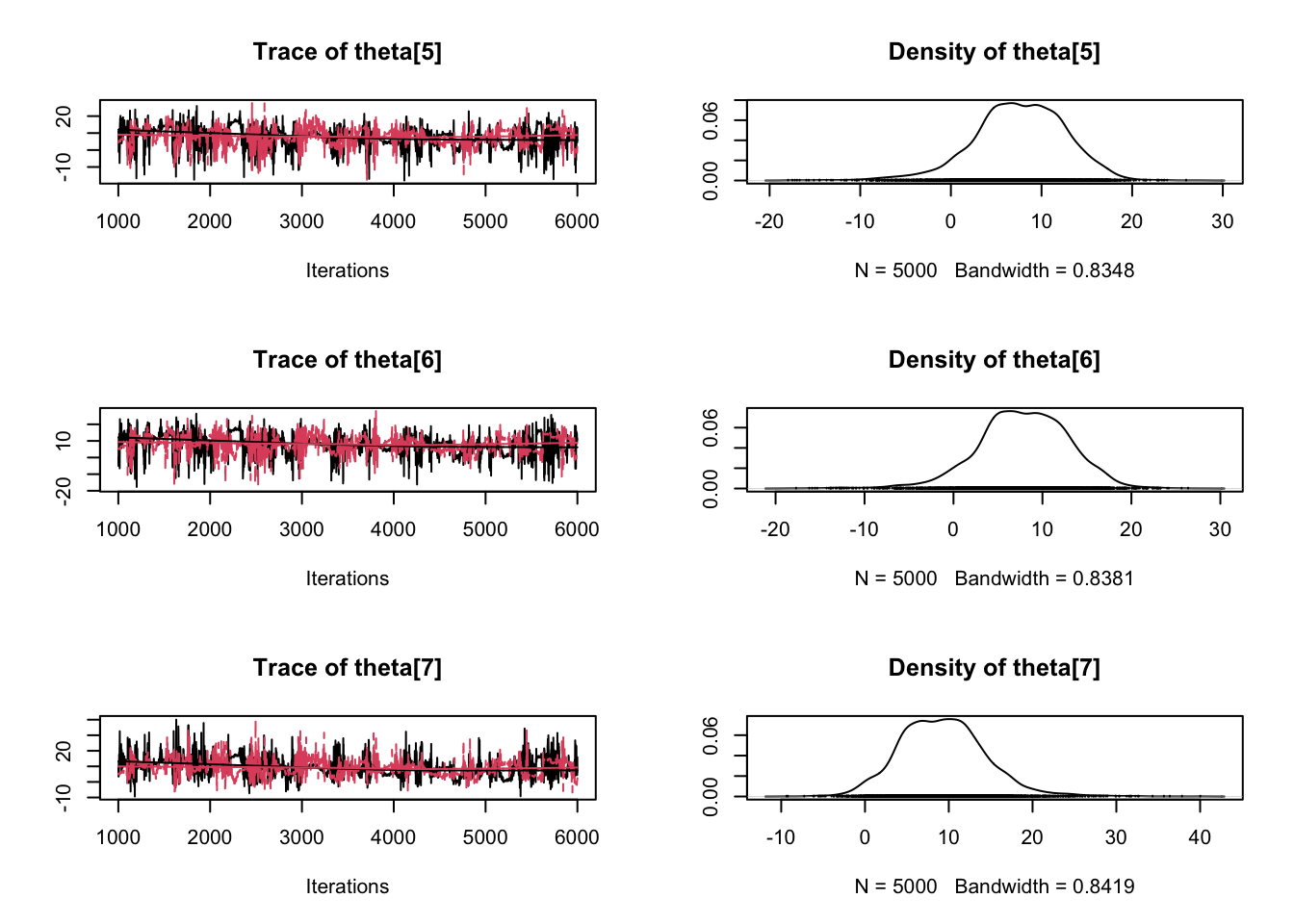
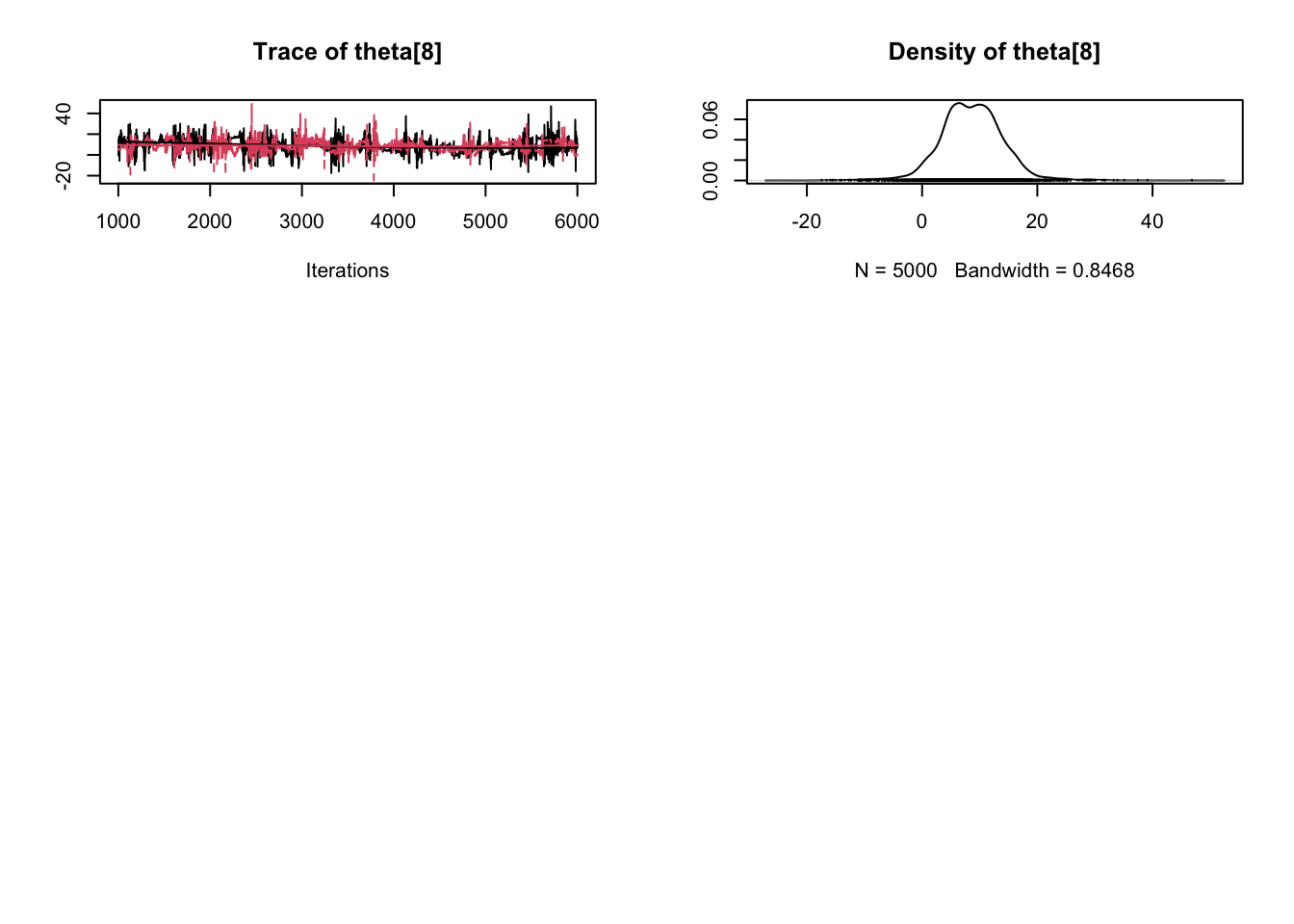
Enjoy Reading This Article?
Here are some more articles you might like to read next: